As COVID-19 ravaged the planet, a small group of investigators at the LSI pivoted their research, and began working at full tilt to identify ways to cripple the SARS-CoV-2 virus.
Biochemistry and Molecular Biology researcher Dr. Natalie Strynadka’s lab was one of four at the Institute to receive support in an early round of federal funding aimed at rapid response to the urgent need for treatment. As the director of the High Resolution Macromolecular Electron Microscopy facility at the Institute, where her research includes a focus on the structure of inhibitors that block antibiotic-resistance, she set her sights on finding a way to block replication of SARS-CoV-2.
Her team’s target was Main Protease (Mpro), an enzyme produced by the virus.
When SARS-CoV-2 infects cells, it starts producing new viral proteins to make more viruses. Initially, a long chain of “polyproteins” is produced from the viral genome, which need to be cut (cleaved) off into individual functional units. In the absence of this essential processing by proteases, including Mpro, the virus cannot replicate. This makes proteases an attractive target for the rapid development of antiviral drugs. Proven drug therapies for other global viral pathogens, such as HIV, also target central proteases.
 In a paper published in Nature Communications, the team revealed how Mpro binds the protein it cleaves (its substrate) to provide important clues as to how it works and how best to design drugs to block its action. This information could also be used to test whether existing antiviral drugs can be repurposed to target Mpro in SARS-CoV-2. Atomic structures and drug discovery typically go hand in hand – particularly for designing enzyme inhibitors.
In a paper published in Nature Communications, the team revealed how Mpro binds the protein it cleaves (its substrate) to provide important clues as to how it works and how best to design drugs to block its action. This information could also be used to test whether existing antiviral drugs can be repurposed to target Mpro in SARS-CoV-2. Atomic structures and drug discovery typically go hand in hand – particularly for designing enzyme inhibitors.
The researchers used X-ray crystallography to capture the atomic structures of Mpro “in action” – bound to substrate — at different stages of the interaction. This is very challenging, because the interactions are very short-lived. Historically, scientists have had little success in capturing these central complexes at the atomic level.
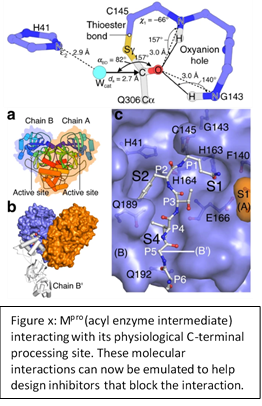
The mechanistic snapshots of Mpro (Figure x) are a major contribution toward understanding how the enzyme works. This is the first time substrate-bound acyl-enzyme intermediate structures have been captured for any Mpro protein, in SARS, MERS or other coronaviruses, and the first snapshot of any cysteine protease.
These atomic blueprints, including ordered water molecules, provide valuable new information for the design of new mechanism-based antivirals to block the devastating effects of COVID-19. Many of the rapidly evolving drug leads targeting SARS-CoV-2 Mpro, whether novel or repurposed, can be readily assessed and optimized using the researchers’ blueprint of the substrate bound complex. The structures also provide a more physiological starting point for future small molecule docking studies being pursued by the Strynadka Lab, as well as many around the globe.
Read the paper
Crystallographic structure of wild-type SARS-CoV-2 main protease acyl-enzyme intermediate with physiological C-terminal autoprocessing site
Jaeyong Lee, Liam J. Worrall, Marija Vuckovic, Federico I. Rosell, Francesco Gentile, Anh-Tien Ton, Nathanael A. Caveney, Fuqiang Ban, Artem Cherkasov, Mark Paetzel & Natalie C. J. Strynadka
Nature Communications volume 11, Article number: 5877 (2020)
