New research from the Lorincz lab makes a significant contribution to answering one of the lingering epigenetic mysteries in developmental biology: how can sperm and eggs of the same species have DNA methylation profiles so different from one another?
In a study recently published in Nature Genetics the researchers have explained the underlying basis of “a significant fraction” of sex-based DNA methylation differences, demonstrating that male and female germ cells use related but distinct histone methyltransferases to direct DNA methylation across the genome.
“Intriguingly, only 40 percent of the genome is methylated in mouse oocytes (eggs), whereas over 80 percent of the genome is methylated in mature sperm,” says First Author Kenjiro Shirane, who recently completed his Banting postdoctoral fellowship at the LSI. “The molecular basis of these sex-based differences in de novo DNA methylation (DNAme) remains an enigma. But in this study, using conditional knockout mice and state-of-the-art sequencing technology, we reveal that distinct histone H3 lysine 36 (H3K36) methyltransferases (KMTases) play instructive roles in de novo DNAme in mouse oocytes and sperm.”
“In essence,” says Senior Author Dr. Matthew Lorincz “this finding is an example of the role of epigenetic cofactors in developmental biology, specifically of the germline, with wide-ranging relevance to medicine and our understanding of the molecular basis of diseases, including overgrowth disorders and cancers, where the same genes we studied are mutated. In other words, there are shared gene regulatory principles at play across development and disease.”
Fundamental to de novo DNAme in eggs, SETD2 is dispensable for methylation in sperm
 Lorincz, Shirane and their collaborators are working towards understanding the interplay between histone modifications and DNAme in mammalian embryos, and germline development. Working with colleagues at Tsinghua University, the team showed in an article published in Nature Genetics in 2019 that SETD2, the lysine methyltransferase responsible for trimethylation of H3K36 (H3K36me3) is required for guiding de novo DNAme by DNMT3A/DNMT3L in mouse oocytes. But the role of histone marks in guiding DNAme in the male germline had, until now, remained a mystery.
Lorincz, Shirane and their collaborators are working towards understanding the interplay between histone modifications and DNAme in mammalian embryos, and germline development. Working with colleagues at Tsinghua University, the team showed in an article published in Nature Genetics in 2019 that SETD2, the lysine methyltransferase responsible for trimethylation of H3K36 (H3K36me3) is required for guiding de novo DNAme by DNMT3A/DNMT3L in mouse oocytes. But the role of histone marks in guiding DNAme in the male germline had, until now, remained a mystery.
“Surprisingly, we found in this new study that SETD2 in sperm is dispensable for de novo DNAme,” says Shirane, now an assistant professor at Kyushu University in Japan. “We went on to profile several other histone modifications and found that H3K36me2 deposited by NSD1 plays an instructive role for de novo DNAme in the male germline.”
Over the last ten years, the Lorincz lab has been studying these proteins and their role in chromatin modification. “NSD1 is one of the many enzymes that add covalent modifications, including methylation, to core histones, which in turn impacts gene expression,” says Lorincz. “We have been trying to understand what the consequences of the lack or addition of those moieties to chromatin are.”
There were dozens of methyltransferases to consider, but NSD1 caught Lorincz’s and his coauthors’ attention after Rosanna Weksberg, a colleague at the University of Toronto, found a broad decrease in DNA methylation in the peripheral blood of her patients with the overgrowth disorder Sotos Syndrome. Children with Sotos, who have loss-of-function mutations on one NSD1 allele, grow rapidly in their first few years of life, and have distinctive facial features, large hands and feet, as well as cognitive and motor delays, speech impairments and low muscle tone.
“We knew of biochemical evidence from embryonic stem cells that trimethylation of histone H3 on lysine 36 is mediated by SETD2, a relative of NSD1,” says Lorincz. “These enzymes are encoded by different genes, and SETD2 is required for recruiting DNA methylation machinery, specifically in the bodies of transcribed genes.
“For some peculiar reason, active genes are DNA methylated in the gene body,” he adds. “Now, if we look in oocytes – as we did in last year’s Nature Genetics paper – the DNA methylation beautifully corresponds to regions that are transcribed. You could almost predict ahead of time, based on biochemical evidence that simply based on the distribution of DNA methylation in oocytes it would likely be SETD2, because this is the only enzyme that adds three methyl groups to lysine 36. But that left the mystery of why does DNA methylation in sperm cover so much more of the genome than in eggs? That’s been an open question for many years.
“DNA methylation is this well-known epigenetic mark, yet there is this really striking difference in methylation between the gametes – a sexually dimorphic pattern that we knew could not be dependent on SETD2 alone in the male germline. Otherwise, you would see a pattern like you see in the female germline, restricted to gene bodies. However, over 80 percent of the genome in sperm are DNA methylated.”
We knew that there must be some other player – most likely a different histone methyltransferase – involved.
As shown in the blood samples from children with Sotos Syndrome, it was possible that an alternative K36 methyltransferase might impact DNA methylation. “Confirming that NSD1 was expressed in male germ cells – we then had the hunch that if we knocked out NSD1 in the male germline – as we had done for SETD2 in female germ cells – that we might see a consequence for DNA methylation in sperm,” says Lorincz.
Knocking out NSD1 in mice is lethal to the embryo, so exploring this hypothesis required a particular conditional knockout mouse, which fortunately had already been generated by colleagues in France. Conditional knockouts allow researchers to delete a gene in a tissue of interest, while leaving it present in the rest of the mouse embryo.
“That’s what we did here, so we could delete NSD1 only in male germ cells,” says Lorincz. “There’s only on the order of a few hundred, or a few thousand germ cells, at the stages we’re studying. So, we used a ultra-low input method for taking snapshots of genome-wide histone modifications called “uli-chip,” which we had previously developed in the lab, that is now widely used in the field. This allowed us to analyze histone modifications with DNA from a few hundred cells, rather than millions, which used to be a limitation. We couldn’t have done these experiments without a low cell input approach.
“Once we had these NSD1 knock out mice, we were off to the races to study the consequences of deletion of this gene in the male germline,” continues Lorincz. “Lysine 36 of Histone H3 is just one residue, one amino acid on a histone of which many of the lysines can be methylated. There are four known enzymes in the mouse or human genome that exclusively methylate Lysine 36 on Histone H3.
An educated guess pays off
“We took an educated guess that NSD1 may be the relevant player in male germ cells, and in much of the genome, and indeed, that turned out to be the case. There was this profound loss of DNA methylation in the NSD1 knockout male germline.”
Circling back to widespread methylation deficits seen in the blood of people with Sotos Syndrome, Lorincz says individuals with the syndrome do express NSD1, but they only have one functional copy of the gene. With only one good copy of this gene in a human, you already have an effect on DNA methylation.
“One of the next set of experiments will be to look into the role of NSD2, a paralog of NSD1, because not all DNA methylation was lost in the NSD1 knockout,” he adds. “Some regions in the male germ cells were still DNA methylated, even in the absence of NSD1, and we suspect that these regions are probably still dependent on K36 methylation, but on a different methyltransferase that deposits that K36 methylation.”
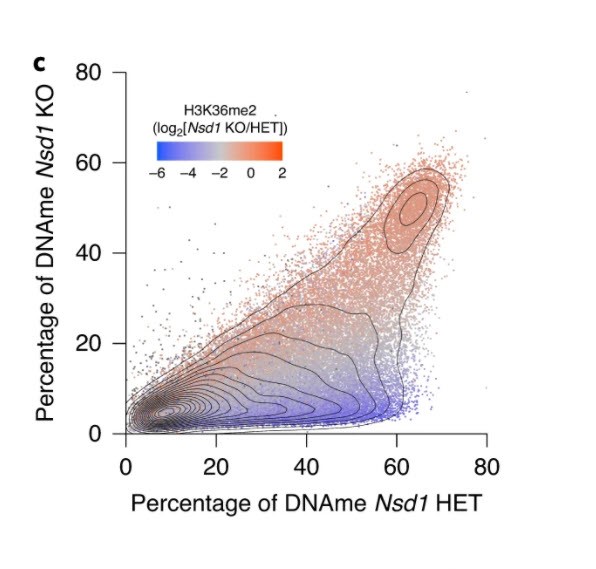 In figure 3, panel C, “the results are as clear as day. The data shows that some regions of the genome are not sensitive to loss of NSD1- the red data points. They remain marked by H3K36me2 and are DNA methylated. In contrast the regions that lose DNA methylation in the absence of NSD1 also lose H3K36me2 in these cells. Importantly, many genes that show such loss are important for male germ cell development.”
In figure 3, panel C, “the results are as clear as day. The data shows that some regions of the genome are not sensitive to loss of NSD1- the red data points. They remain marked by H3K36me2 and are DNA methylated. In contrast the regions that lose DNA methylation in the absence of NSD1 also lose H3K36me2 in these cells. Importantly, many genes that show such loss are important for male germ cell development.”
“So, we get this important phenotype, with now the added complexity that another epigenetic mark, H3K27 trimethylation, (H3K27me3) which is deposited by yet another histone methyltransferase called EZH2, is increased at a subset of the regions that lose H3K36me2. That’s a very well-studied enzyme and has a history here at UBC, because Bill Gibson in the Department of Medical Genetics identified mutations in EZH2 as the cause of Weaver Syndrome. Interestingly enough, it shares many similarities to Sotos Syndrome, including overgrowth. Surprisingly, this gain of H3K27me3 in regions losing H3K36me2 and DNA methylation leads to transcriptional silencing of nearby genes, which we believe is responsible for the ultimate failure to generate sperm in male germ cells deficient in NSD1.”
Figure 6 from the paper explains this scenario in cartoon form.

The orange colour on the left shows what happens in oocytes, and on the right is what’s happening in the male germline. At the top on the right, the orange represents the histone H3K36 methylation being deposited by NSD1.
DNMT3A in the second row is the DNA methyltransferase that is recruited to specific regions of the genome, by the presence of the H3K36 methylation. It then methylates the DNA, as illustrated by the lollipops, going from white to black, showing that “de novo” DNA methylation.
H3K27 methylation levels are actually increased when H3K36 methylation and DNA methylation levels go down. This increase in H3K27me3 is associated with downregulation of genes that would otherwise be expressed. So, the K36 methylation is actually counteracting H3K27 methylation mediated by EZH2.
A fascination with the most basic aspects of development – and their implications
“I would say my main interest is in the fundamentals of developmental biology. What makes the system tick in terms of the interplay between epigenetic marks and gene expression,” says Lorincz, when asked what he wished to solve or address in looking into this long-standing mystery. “What are the features of chromatin? The proteins that regulate chromatin – what role do they play in directing developmental processes, like, the development of the gametes or early embryo?
“While my interests are really focused on understanding how this metaphorical mousetrap is built, it turns out that the proteins involved in normal development also play very important roles in diseases such as overgrowth syndromes and cancer.
“So, arising from basic research questions, potential great insights can come in our understanding of disease — because you can’t manipulate cells and study rare cells in the way you can using an animal model, like the mouse, if you’re studying human tissues. Furthermore, interesting parallels emerge when comparing results from mouse models to those from cancer cells. For example, a group at McGill studying head and neck cancers has found that the subset harboring mutations in NSD1 also have broadly lower levels of DNA methylation.
“NSD1 pokes its head both in cancer and in developmental disorders,” reflects Lorincz, “with the consistent observation being that this histone mark has this important role in directing DNA methylation, and the important point is that there are downstream consequences for gene expression when these epigenetic marks are disrupted.”
NSD2 will be the next focus
“The finding in this study will enhance our understanding of sexually dimorphic patterns of gene regulation in oocytes and sperm,” concludes first-author Dr. Kenjiro Shirane. Many questions, including why oocytes and sperm use distinct enzymes to direct de novo DNAme, remain to be answered. As male mice deficient in germline NSD1 show infertility, NSD1 and H3K36me2 deposited by this enzyme could potentially be used as biomarkers and clinical intervention of human infertility.”
NSD2 is of interest, as it too is responsible for a developmental syndrome, known as Wolf-Hirschhorn — that can also be mutated in cancer. “Interestingly enough, contrary to Sotos Syndrome, people with Wolf-Hirschhorn syndrome experience delayed growth and development. But what role it plays in germline, and embryonic development has not been explored,” says Lorincz. “It’s possible that knock out of NSD2 in the male germline, unlike NSD1, will have no effect on fertility. Nevertheless, DNA methylation may be reduced in certain genomic regions, but these may be distinct from those that NSD1 influences.”
“Long-term,” reflects Lorincz, “we’re interested in trying to further understand the role that histone modifying enzymes, like NSD1, or NSD2, or the DNA methyltransferases, have in gene regulation, with the ultimate goal of deciphering the complex cross-talk between these epigenetic marks. That’s really the most important thing, because you can disrupt a specific histone methyltransferase, and that will not only impact the histone mark that methyltransferase deposits, but also secondarily, disrupts not only the normal distribution of DNA methylation but also that of other histone marks, like Lysine 27 methylation, which are deposited by totally different enzymes. It’s almost like there’s this domino effect. You affect one, and it has consequences that you wouldn’t necessarily a priori be able to predict. You have to do the experiment and see what the consequences are.”
Read the paper
Shirane, K., Miura, F., Ito, T., Lorincz, M.C. NSD1-deposited H3K36me2 de novo methylation in the mouse male germline and counteracts Polycomb-associated silencing. Nat. Genet. 52, 1088-1098 (2020).
This research was funded by
The Platform Project for Supporting Drug Discovery and Life Science Research (Basis for Supporting Innovative Drug Discovery and Life Science Research (BINDS) from AMED (Japan Agency for Medical Research and Development). M.C.L. is supported by CIHR grants, and K.S. was a recipient of a Uehara Memorial Foundation postdoctoral fellowship and a CIHR Banting postdoctoral fellowship.
