Molecular Mechanisms of Autophagy Regulation
Neurodegenerative diseases (ND), including Alzheimer’s disease (AD), Parkinson’s disease (PD), Huntington’s disease (HD), and Amyotrophic Lateral Sclerosis (ALS), are leading causes of disability in Canada, affecting over 800,000 Canadians and incur >$8 billion annually in healthcare spending. With the aging Canadian population, the incidence of ND is projected to rise dramatically over the next decade, and there is thus an urgent need to find effective treatment(s) for these currently incurable human pathologies. This is the important area that the LSI’s Dr. Calvin Yip will be researching using his recently awarded 2016 CIHR Foundation Grant.
Research Summary
A common pathological hallmark of ND is the cytoplasmic accumulation of protein aggregates that are positively associated with neuronal toxicity. Although harmful genetic mutations can render certain proteins more prone to misfolding and aggregation, recent studies suggest that neurons normally have the capability to remove these mutant proteins and their aggregates using the autophagy degradation system. Mounting evidence pointed to impairment of the autophagy system as a major driving force behind the toxic buildup of protein aggregates in ND. In particular, two landmark animal-based studies demonstrated that central nervous system specific ablation of autophagy causes aggregate accumulation and severe ND symptoms even in the absence of harmful mutations. These studies not only confirmed the link between autophagy and ND but also emphasized the importance of understanding how autophagy is regulated in normal cells and dysregulated in the disease state in developing therapeutic strategies for ND.
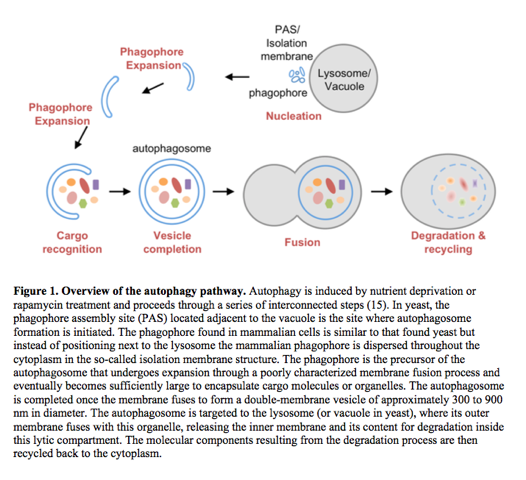
Autophagy degradation proceeds in multiple steps to sequester cytoplasmic cargo into an autophagosomal vesicle and to deliver it to the lysosome for breakdown (see Fig. 1 below), but the molecular mechanisms underlying these processes are poorly defined. The discovery of the ~30 ATG (autophagy related) genes by yeast genetic screens and the recent proteomic mapping of the ~400-protein human autophagy network revealed the identity and the modular nature of the core autophagy machinery of autophagy.
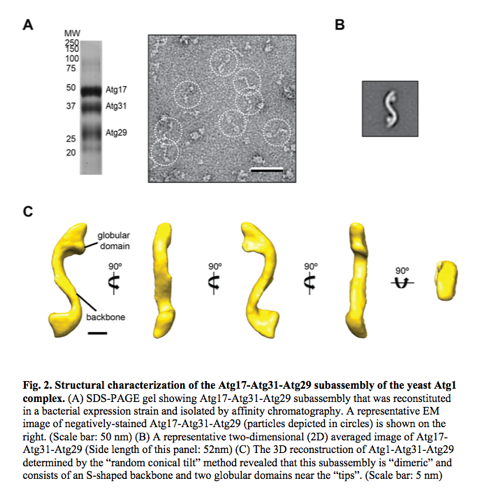
Yip believes that gaining a structural perspective of the various autophagy modules will provide mechanistic insights into fundamental autophagy regulatory mechanisms critical to understanding the molecular etiologies of ND. His lab’s recent studies on the Atg17-Atg31-Atg29 subassembly of the autophagy signaling module provided proof-of-concept for their single-particle EM molecular imaging approach (see Fig. 2 below) and placed them in a unique position to expand our structure-function analyses to other components of this module as well as to other key modules.